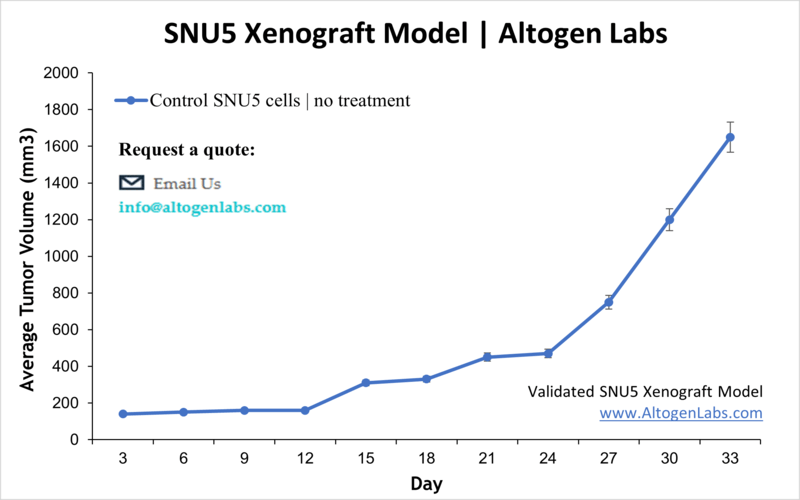
SNU5 Xenograft Model (Standard CDX)
SNU-5 is a human gastric cancer cell line that was established from a patient with a poorly differentiated adenocarcinoma of the stomach. Gastric adenocarcinoma is a common type of stomach cancer that arises from the glandular cells lining the stomach. Over 200,000 cases of gastric cancer occur annually in the United States. This disease often has little to no early-stage symptoms and the most common cause of gastric cancer is infection with the bacterium Helicobacter pylori. The SNU5 cell line was isolated by J. Park in 1987 from the metastatic ascites of a female Asian 33 year old patient with gastric carcinoma. SNU5 cells have since been used as a colon cancer model in many studies. A 2012 Clinical Cancer Research study by Kanjoormana et al. used the SNU5 xenograft model to characterize the ability of γ-tocotrienol to sensitize tumors to capecitabine. Results demonstrated that combination treatment with γ-tocotrienol and capecitabine led to synergistic NFΚ-B-mediated inhibition of proliferation and tumor growth marked by the suppression of markers of invasion, proliferation, metastasis and angiogenesis. Fujian et al. released a Drug Design, Development and Therapy article in 2015 using the SNU5 xenograft model to characterize a mechanism of MET resistance to PHA665752, a MET inhibitor. Results showed that PI3K p110α overexpression is a biomarker for the tyrosine kinase inhibitor resistance, supporting PI3K p110α as a novel potential target for gastrinoma. Lastly, in 2014 Liu et al. published a study in Tumour Biology using SNU5 subcutaneous xenografts to characterize the anticancer effects of oridonin, a diterpenoid compound isolated from Rabdosia rubescens. SNU5 was chosen for the c-Met gene amplification; results demonstrated that oridonin treatment was well tolerated and anti-tumor activity was dose dependent and manifested as reduction in microvessel density (anti-angiogenesis as noted by CD31 levels), suppressed tumor proliferation (measured by Ki67) and growth inhibition. The SNU5 cell line is used to create the CDX (Cell Line Derived Xenograft) SNU5 xenograft mouse model. The SNU5 xenograft model has been used in gastric cancer studies including combination therapies and MET resistance.
Basic Study Design
- SNU5 cells are maintained in exponential growth phase under aseptic conditions.
- Cells are trypsinized and cell count viability is determined using a trypan blue exclusion assay (99% of cell viability required). SNU5 cell suspension is adjusted to appropriate density.
- Each mouse is singly subcutaneously injected into the right flank with 10e6 cells in 100-200 µL of a Matrigel-SNU5 cell suspension.
- The injection sites are palpated up to three times weekly until tumors are established to an average size of 90-140 cubic mm as measured by digital calipers.
- Animals are randomized into treatment groups. Administration of test compound is performed according to the pre-established treatment schedule.
- Mice weights are measured and recorded 2-3 times weekly; tumors are measured and recorded daily.
- End of study is reached when tumor size reaches 2,000 cubic mm or the predetermined size limit per approved IACUC protocol.
- Final necropsy and tissue collections are performed for appropriate downstream analysis. Tumors are excised, weighed and documented by digital imaging. Tumors and tissues can be stabilized in RNA-later reagent, snap frozen in LN2 or prepared for histology.
